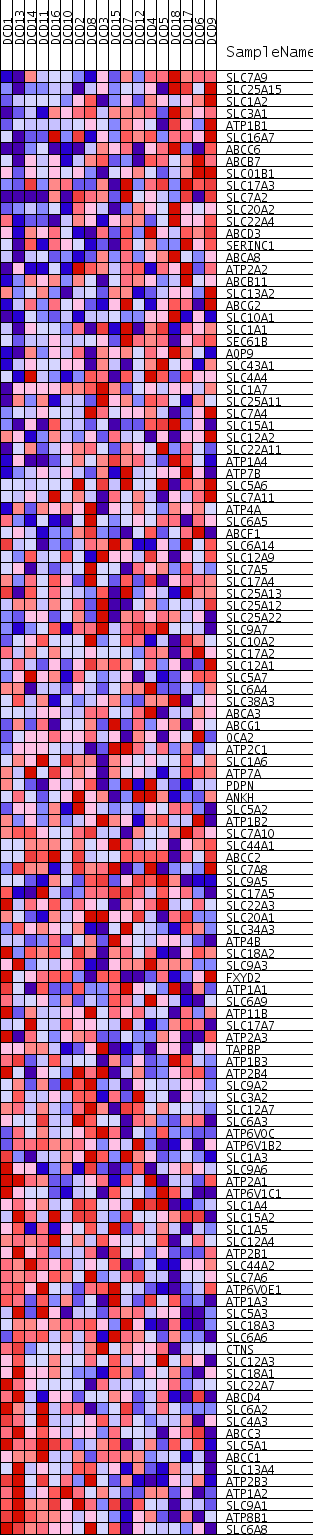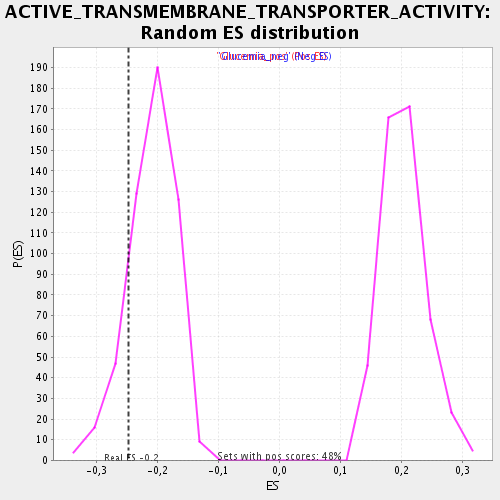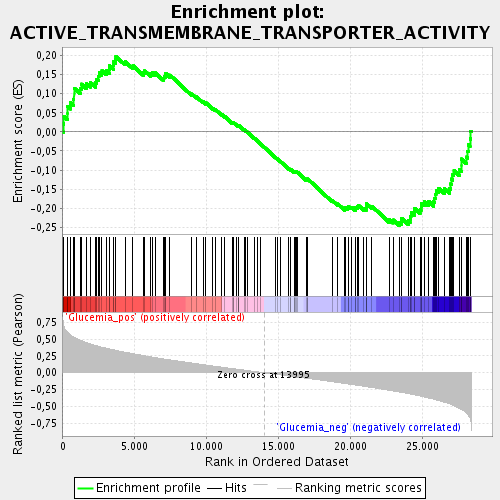
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | DCD_collapsed_to_symbols.DCD.cls#Glucemia |
| Phenotype | DCD.cls#Glucemia |
| Upregulated in class | Glucemia_neg |
| GeneSet | ACTIVE_TRANSMEMBRANE_TRANSPORTER_ACTIVITY |
| Enrichment Score (ES) | -0.24657339 |
| Normalized Enrichment Score (NES) | -1.1829715 |
| Nominal p-value | 0.15163147 |
| FDR q-value | 0.77359414 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC7A9 | 15 | 0.744 | 0.0216 | No | ||
| 2 | SLC25A15 | 65 | 0.690 | 0.0403 | No | ||
| 3 | SLC1A2 | 322 | 0.602 | 0.0491 | No | ||
| 4 | SLC3A1 | 327 | 0.601 | 0.0668 | No | ||
| 5 | ATP1B1 | 523 | 0.558 | 0.0765 | No | ||
| 6 | SLC16A7 | 725 | 0.532 | 0.0852 | No | ||
| 7 | ABCC6 | 768 | 0.528 | 0.0994 | No | ||
| 8 | ABCB7 | 769 | 0.528 | 0.1151 | No | ||
| 9 | SLCO1B1 | 1196 | 0.486 | 0.1145 | No | ||
| 10 | SLC17A3 | 1282 | 0.477 | 0.1257 | No | ||
| 11 | SLC7A2 | 1648 | 0.447 | 0.1261 | No | ||
| 12 | SLC20A2 | 1889 | 0.431 | 0.1304 | No | ||
| 13 | SLC22A4 | 2256 | 0.407 | 0.1295 | No | ||
| 14 | ABCD3 | 2350 | 0.401 | 0.1382 | No | ||
| 15 | SERINC1 | 2461 | 0.394 | 0.1460 | No | ||
| 16 | ABCA8 | 2512 | 0.391 | 0.1558 | No | ||
| 17 | ATP2A2 | 2714 | 0.380 | 0.1600 | No | ||
| 18 | ABCB11 | 3020 | 0.365 | 0.1601 | No | ||
| 19 | SLC13A2 | 3217 | 0.356 | 0.1638 | No | ||
| 20 | ABCG2 | 3239 | 0.355 | 0.1736 | No | ||
| 21 | SLC10A1 | 3505 | 0.342 | 0.1744 | No | ||
| 22 | SLC1A1 | 3529 | 0.341 | 0.1837 | No | ||
| 23 | SEC61B | 3642 | 0.335 | 0.1897 | No | ||
| 24 | AQP9 | 3684 | 0.333 | 0.1981 | No | ||
| 25 | SLC43A1 | 4322 | 0.306 | 0.1847 | No | ||
| 26 | SLC4A4 | 4872 | 0.284 | 0.1738 | No | ||
| 27 | SLC1A7 | 5604 | 0.257 | 0.1556 | No | ||
| 28 | SLC25A11 | 5693 | 0.253 | 0.1600 | No | ||
| 29 | SLC7A4 | 6086 | 0.239 | 0.1532 | No | ||
| 30 | SLC15A1 | 6208 | 0.234 | 0.1559 | No | ||
| 31 | SLC12A2 | 6429 | 0.226 | 0.1548 | No | ||
| 32 | SLC22A11 | 7009 | 0.206 | 0.1405 | No | ||
| 33 | ATP1A4 | 7032 | 0.205 | 0.1458 | No | ||
| 34 | ATP7B | 7111 | 0.202 | 0.1490 | No | ||
| 35 | SLC5A6 | 7152 | 0.201 | 0.1536 | No | ||
| 36 | SLC7A11 | 7436 | 0.192 | 0.1493 | No | ||
| 37 | ATP4A | 8964 | 0.145 | 0.0996 | No | ||
| 38 | SLC6A5 | 9272 | 0.136 | 0.0928 | No | ||
| 39 | ABCF1 | 9747 | 0.122 | 0.0797 | No | ||
| 40 | SLC6A14 | 9943 | 0.116 | 0.0762 | No | ||
| 41 | SLC12A9 | 10436 | 0.100 | 0.0618 | No | ||
| 42 | SLC7A5 | 10593 | 0.096 | 0.0592 | No | ||
| 43 | SLC17A4 | 11019 | 0.084 | 0.0466 | No | ||
| 44 | SLC25A13 | 11268 | 0.076 | 0.0401 | No | ||
| 45 | SLC25A12 | 11771 | 0.062 | 0.0242 | No | ||
| 46 | SLC25A22 | 11856 | 0.060 | 0.0230 | No | ||
| 47 | SLC9A7 | 11881 | 0.059 | 0.0240 | No | ||
| 48 | SLC10A2 | 12062 | 0.054 | 0.0192 | No | ||
| 49 | SLC17A2 | 12185 | 0.050 | 0.0164 | No | ||
| 50 | SLC12A1 | 12217 | 0.049 | 0.0167 | No | ||
| 51 | SLC5A7 | 12245 | 0.049 | 0.0172 | No | ||
| 52 | SLC6A4 | 12597 | 0.039 | 0.0060 | No | ||
| 53 | SLC38A3 | 12713 | 0.036 | 0.0030 | No | ||
| 54 | ABCA3 | 12870 | 0.032 | -0.0016 | No | ||
| 55 | ABCG1 | 13322 | 0.019 | -0.0169 | No | ||
| 56 | OCA2 | 13528 | 0.013 | -0.0238 | No | ||
| 57 | ATP2C1 | 13724 | 0.008 | -0.0304 | No | ||
| 58 | SLC1A6 | 14762 | -0.022 | -0.0664 | No | ||
| 59 | ATP7A | 14910 | -0.026 | -0.0708 | No | ||
| 60 | PDPN | 15114 | -0.031 | -0.0771 | No | ||
| 61 | ANKH | 15719 | -0.048 | -0.0970 | No | ||
| 62 | SLC5A2 | 15848 | -0.052 | -0.1000 | No | ||
| 63 | ATP1B2 | 15856 | -0.052 | -0.0986 | No | ||
| 64 | SLC7A10 | 16077 | -0.058 | -0.1047 | No | ||
| 65 | SLC44A1 | 16127 | -0.060 | -0.1046 | No | ||
| 66 | ABCC2 | 16156 | -0.060 | -0.1038 | No | ||
| 67 | SLC7A8 | 16232 | -0.063 | -0.1046 | No | ||
| 68 | SLC9A5 | 16304 | -0.064 | -0.1052 | No | ||
| 69 | SLC17A5 | 16933 | -0.081 | -0.1250 | No | ||
| 70 | SLC22A3 | 16979 | -0.083 | -0.1241 | No | ||
| 71 | SLC20A1 | 17026 | -0.084 | -0.1233 | No | ||
| 72 | SLC34A3 | 18729 | -0.132 | -0.1794 | No | ||
| 73 | ATP4B | 19117 | -0.145 | -0.1888 | No | ||
| 74 | SLC18A2 | 19593 | -0.159 | -0.2009 | No | ||
| 75 | SLC9A3 | 19661 | -0.161 | -0.1984 | No | ||
| 76 | FXYD2 | 19831 | -0.166 | -0.1995 | No | ||
| 77 | ATP1A1 | 19844 | -0.167 | -0.1949 | No | ||
| 78 | SLC6A9 | 20050 | -0.173 | -0.1970 | No | ||
| 79 | ATP11B | 20341 | -0.181 | -0.2019 | No | ||
| 80 | SLC17A7 | 20371 | -0.182 | -0.1975 | No | ||
| 81 | ATP2A3 | 20476 | -0.186 | -0.1957 | No | ||
| 82 | TAPBP | 20576 | -0.189 | -0.1935 | No | ||
| 83 | ATP1B3 | 20927 | -0.201 | -0.1999 | No | ||
| 84 | ATP2B4 | 21089 | -0.206 | -0.1995 | No | ||
| 85 | SLC9A2 | 21105 | -0.207 | -0.1939 | No | ||
| 86 | SLC3A2 | 21109 | -0.207 | -0.1878 | No | ||
| 87 | SLC12A7 | 21482 | -0.220 | -0.1945 | No | ||
| 88 | SLC6A3 | 22691 | -0.260 | -0.2294 | No | ||
| 89 | ATP6V0C | 22962 | -0.270 | -0.2310 | No | ||
| 90 | ATP6V1B2 | 23405 | -0.287 | -0.2380 | Yes | ||
| 91 | SLC1A3 | 23537 | -0.292 | -0.2340 | Yes | ||
| 92 | SLC9A6 | 23565 | -0.293 | -0.2263 | Yes | ||
| 93 | ATP2A1 | 24023 | -0.310 | -0.2332 | Yes | ||
| 94 | ATP6V1C1 | 24150 | -0.316 | -0.2283 | Yes | ||
| 95 | SLC1A4 | 24201 | -0.317 | -0.2206 | Yes | ||
| 96 | SLC15A2 | 24217 | -0.318 | -0.2117 | Yes | ||
| 97 | SLC1A5 | 24433 | -0.327 | -0.2096 | Yes | ||
| 98 | SLC12A4 | 24466 | -0.329 | -0.2009 | Yes | ||
| 99 | ATP2B1 | 24845 | -0.344 | -0.2041 | Yes | ||
| 100 | SLC44A2 | 24915 | -0.347 | -0.1962 | Yes | ||
| 101 | SLC7A6 | 24940 | -0.349 | -0.1867 | Yes | ||
| 102 | ATP6V0E1 | 25129 | -0.359 | -0.1827 | Yes | ||
| 103 | ATP1A3 | 25429 | -0.375 | -0.1821 | Yes | ||
| 104 | SLC5A3 | 25797 | -0.394 | -0.1834 | Yes | ||
| 105 | SLC18A3 | 25859 | -0.397 | -0.1738 | Yes | ||
| 106 | SLC6A6 | 25921 | -0.400 | -0.1640 | Yes | ||
| 107 | CTNS | 25956 | -0.403 | -0.1533 | Yes | ||
| 108 | SLC12A3 | 26140 | -0.414 | -0.1474 | Yes | ||
| 109 | SLC18A1 | 26561 | -0.440 | -0.1492 | Yes | ||
| 110 | SLC22A7 | 26896 | -0.463 | -0.1473 | Yes | ||
| 111 | ABCD4 | 26988 | -0.471 | -0.1365 | Yes | ||
| 112 | SLC6A2 | 27017 | -0.474 | -0.1234 | Yes | ||
| 113 | SLC4A3 | 27121 | -0.482 | -0.1127 | Yes | ||
| 114 | ABCC3 | 27204 | -0.490 | -0.1011 | Yes | ||
| 115 | SLC5A1 | 27580 | -0.530 | -0.0986 | Yes | ||
| 116 | ABCC1 | 27710 | -0.543 | -0.0870 | Yes | ||
| 117 | SLC13A4 | 27712 | -0.543 | -0.0710 | Yes | ||
| 118 | ATP2B3 | 28069 | -0.598 | -0.0658 | Yes | ||
| 119 | ATP1A2 | 28155 | -0.618 | -0.0504 | Yes | ||
| 120 | SLC9A1 | 28212 | -0.634 | -0.0336 | Yes | ||
| 121 | ATP8B1 | 28322 | -0.674 | -0.0174 | Yes | ||
| 122 | SLC6A8 | 28370 | -0.710 | 0.0020 | Yes |